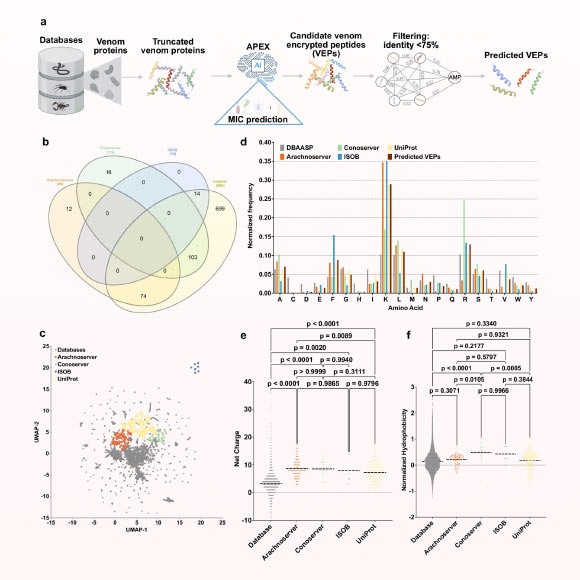
University of Pennsylvania researchers used a deep learning tool named Apex to explore a worldwide venom dataset in search of new antibiotic candidates.
Guan et al. Vococcus is a rich source of previously hidden antibiotic scaffolds, showing that merging experimental validation with extensive computational mining can enhance the search for urgently needed antibiotics. Image credits: Guan et al., doi: 10.1038/s41467-025-60051-6.
The increasing prevalence of antibiotic-resistant pathogens, especially Gram-negative bacteria, underscores the critical demand for new treatments.
Venococcus represents a vast, largely untapped source of bioactive molecules with potential antibacterial properties.
In their recent study, researcher César de La Fuente and his team analyzed a comprehensive database containing 16,123 poison proteins and over 40 million poison-encoded peptides via a vertex deep learning model.
The algorithm successfully pinpointed 386 candidate peptides that differ in structure and function from known antimicrobial peptides.
“These poisons are evolutionary wonders, yet their antibacterial capabilities have not been thoroughly examined,” said Dr. de la Fuente.
“Apex can rapidly explore extensive chemical landscapes, identifying exceptional peptides that combat some of the most stubborn pathogens worldwide.”
From the potential candidates selected by AI, scientists synthesized 58 peptide variants for laboratory assessment.
Remarkably, 53 of these demonstrated efficacy against drug-resistant bacteria such as E. coli and Staphylococcus aureus, at doses safe for human red blood cells.
“By combining computational analysis with traditional laboratory techniques, we achieved one of the most thorough antibiotic studies to date,” noted Dr. Marcelo Torres, co-author of the research.
“The platform has mapped over 2,000 novel antibacterial motifs, enhancing its capacity to eliminate or suppress bacterial growth through short, specific amino acid sequences within proteins or peptides.”
“Our team is now advancing the top peptide candidates towards the development of new antibiotics, optimizing them through medicinal chemistry modulation.”
results will be published in the journal Nature Communications.
____
C. Guan et al. 2025. A global assessment of venom data for antibacterial discovery using artificial intelligence techniques. Nat Commun 16, 6446; doi:10.1038/s41467-025-60051-6
Source: www.sci.news