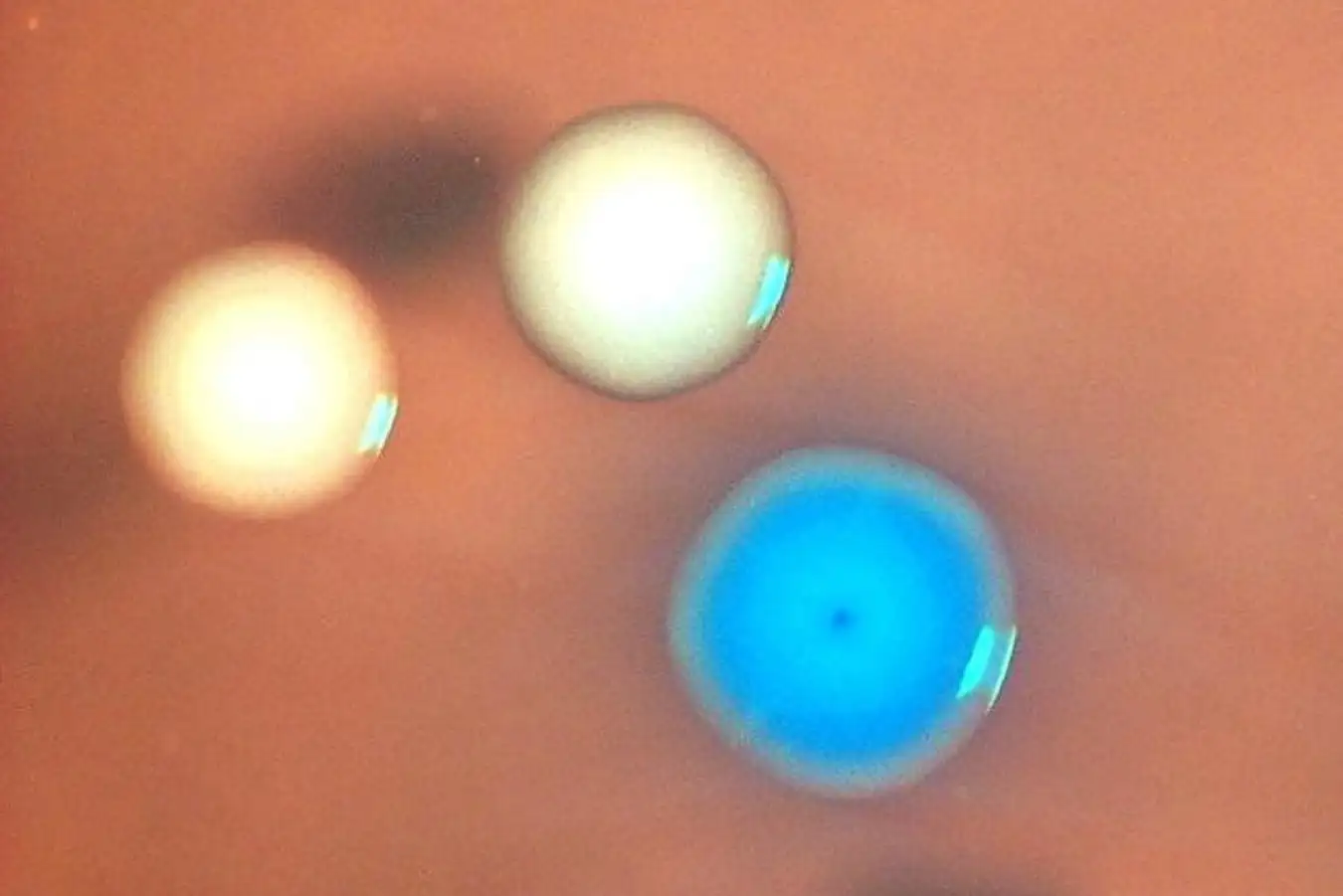
Microscopic View of Bacterial Colonies: Blue Colonies Represent Synthetic Genomes, While White Colonies Show Survivors of Mitomycin C Treatment.
Credit: Nasaira Assad-Garcia
Researchers have successfully developed living synthetic cells by transplanting complete genomes into deceased bacteria, effectively bringing these microorganisms back to life. This groundbreaking advancement has the potential to revolutionize synthetic biology, allowing for the engineering of living organisms to produce sustainable fuels, pharmaceuticals, and novel materials.
Synthetic biology involves modifying biological systems to introduce new functionalities or create entirely new systems. For instance, scientists can rewrite yeast DNA so that these organisms can synthesize desired chemicals. In 2010, groundbreaking work saw researchers synthesizing bacterial genomes and deploying them into living cells, birthing what they termed as the first synthetic cells.
However, challenges arose; determining whether the cells were genuinely driven by the synthetic genome rather than the original was complex. This issue stemmed from bacteria’s ability to absorb external genetic material via horizontal gene transfer.
To overcome this, John Glass and colleagues at the J. Craig Venter Institute (JCVI) in La Jolla, California, first exterminated the host cell—or at least its genome.
The team employed a chemical called mitomycin C, commonly used in chemotherapy to damage DNA, and tested it on simple bacterial cells of mycoplasma capricolum.
The researchers noted, “The cells remain healthy but are unable to reproduce and their genomes are non-functional, leaving them destined for death or already deceased,” according to Zumra Seidel, also from JCVI.
Next, they introduced a synthetic variant of another bacterial genome from Mycoplasma mycoides into the dead cells through whole-genome transplantation.
Surprisingly, some bacteria began to grow and replicate normally, with genetic tests confirming the presence of synthetic genomes. The team proudly claimed to have engineered the first living synthetic bacterial cells derived from non-living components, dubbing them “zombie cells” due to their revival post-mortem.
“Introducing a genome to a cell devoid of one restores its functionality,” explained Glass.
Kate Adamara from the University of Minnesota commended this research as a pivotal technological breakthrough. “They are embedding genomic information into a non-living recipient with no assistance from the host’s repair systems. Essentially, they have revived that cell,” she noted. “An impressive feat!”
Furthermore, it raises questions about the definitions of life and non-life; traditionally defined by metabolism and replication, these traits are barely present in the recipient cells. “What truly constitutes life?” queried Adamara.
Team member Elizabeth Strychalski from the National Institute of Standards and Technology in Gaithersburg, Maryland, expressed hope that this discovery would encourage viewing biology as fluid processes. “By adopting an engineering perspective, we can analyze living systems and identify which processes are essential for our desired outcomes,” she stated.
This technique has thus far only been tested on mycoplasma, yet the researchers believe it serves as a proof of principle and could significantly expedite the creation of synthetic organisms that function as mini-chemical factories to produce drugs or clean up environmental pollutants.
“While we have long had the capability to assemble remarkable lengths of synthetic DNA, we lacked means to make them operational,” Strychalski remarked. “It’s akin to having a script for a Shakespearean play without the ability to perform it.”
Akos Nierges from Harvard Medical School emphasized that this research tackles a vital hurdle in synthetic biology. “This technology may lead to more predictable and reliable methods for genome transfer across various species,” he said.
Transitioning to more complex organisms like yeast and Escherichia coli could pose challenges due to their cell walls. Still, Glass remains optimistic that this technology can succeed with those genomes too.
“If effective in one organism, it’s likely to succeed in others,” he stated, with ongoing investigations into methods to remove and replace cell walls. “Provided appropriate growth conditions, Escherichia coli can regenerate new cell walls,” he added.
Concerns about biological safety in synthetic biology persist. Although the mycoplasma species examined in this study can be pathogens for goats and cattle, Nierges assured there are no anticipated increases in virulence from these modifications.
Strychalski mentioned that existing best practices in laboratories can significantly reduce the risk of pathogen leakage.
Topics:
- Biotechnology /
- Microbiology
Source: www.newscientist.com